Our key publications
S. Robert group members in bold
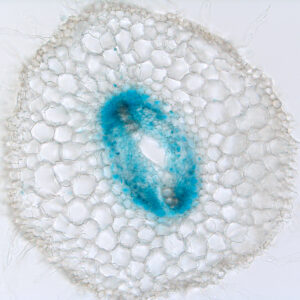
Auxin triggers pectin modification during rootlet emergence in white lupin
Plant J 2022, 112(5):1127-40
Jobert F, Soriano A, Brottier L, Casset C, Divol F, Safran J, Lefebvre V, Pelloux J, Robert S*, Péret B* (*joint corresponding authors)
https://doi.org/10.1111/tpj.15993
-> See our Plant Journal cover here!
-> See also the Research Highlight written about our article!
———
A network of stress-related genes regulates hypocotyl elongation downstream of selective auxin perception
Plant Physiol 2021, 187(1):430-45
Rigal A*, Doyle SM*, Ritter A, Raggi S, Vain T, O’Brien JA, Goossens A, Pauwels L, Robert S (*joint first authors)
https://doi.org/10.1093/plphys/kiab269
-> See our Plant Physiology first author highlight!
———
New fluorescent auxin probes visualize tissue-specific and subcellular distributions of auxin in Arabidopsis
New Phytol 2021, 230:535-549
Pařízková B*, Žukauskaitė A*, Vain T*, Grones P*, Raggi S, Kubeš MF, Kieffer M, Doyle SM, Strnad M, Kepinski S, Napier R, Doležal K, Robert S^, Novák O^ (*joint first authors; ^joint corresponding authors)
https://doi.org/10.1111/nph.17183
———
Fluctuating auxin response gradients determine pavement cell-shape acquisition
Proc Natl Acad Sci USA 2020, 117(27):16027-16034
Grones P, Majda M, Doyle SM, Van Damme D, Robert S
https://doi.org/10.1073/pnas.2007400117
———
A role for the auxin precursor anthranilic acid in root gravitropism via regulation of PIN-FORMED protein polarity and relocalisation in Arabidopsis
New Phytol 2019, 223(3):1420-1432
Doyle SM*, Rigal A*, Grones P, Karady M, Barange DK, Majda M, Pařízková B, Karampelias M, Zwiewka M, Pěnčik A, Almqvist F, Ljung K, Novák O, Robert S (*joint first authors)
https://doi.org/10.1111/nph.15877
———
Mechanical asymmetry of the cell wall predicts changes in pavement cell geometry
Dev Cell 2019, 50(1):9-10
Majda M, Krupinski P, Jönsson H, Hamant O, Robert S
https://doi.org/10.1016/j.devcel.2019.06.002
———
Selective auxin agonists induce specific AUX/IAA protein degradation to modulate plant development
Proc Natl Acad Sci USA 2019, 116(13):6463-6472
Vain T*, Raggi S*, Ferro N, Barange DK, Kieffer M, Ma Q, Doyle SM, Thelander M, Pařízková B, Novák O, Ismail A, Enquist PA, Rigal A, Łangowska M, Ramans Harborough S, Zhang Y, Ljung K, Callis J, Almqvist F, Kepinski S, Estelle M, Pauwels L, Robert S (*joint first authors)
https://doi.org/10.1073/pnas.1809037116
———
Mechanochemical polarization of contiguous cell walls shapes plant pavement cells
Dev Cell 2017, 43(3): 290–304.e4
Majda M, Grones P, Sintorn I-M, Vain T, Milani P, Krupinski P, Zagórska-Marek B, Viotti C, Jönsson H, Mellerowicz EJ, Hamant O, Robert S
https://doi.org/10.1016/j.devcel.2017.10.017
———
2,4-D and IAA amino acid conjugates show distinct metabolism in Arabidopsis
PLoS One 2016, 11(7):e0159269
Eyer L, Vain T, Pařízková B, Oklestkova J, Barbez E, Kozubíková H, Pospíšil T, Wierzbicka R, Kleine-Vehn J, Fránek M, Strnad M, Robert S*, Novak O* (*joint corresponding authors)
https://doi.org/10.1371/journal.pone.0159269
———
An early secretory pathway mediated by GNOM-LIKE 1 and GNOM is essential for basal polarity establishment in Arabidopsis thaliana
Proc Natl Acad Sci USA 2015, 112(7):E806-15
Doyle SM, Haeger A, Vain T, Rigal A, Viotti C, Łangowska M, Ma Q, Friml J, Raikhel NV, Hicks GR, Robert S
https://doi.org/10.1073/pnas.1424856112
———
All SR group publications
S. Robert group members in bold
Guidelines for naming and studying plasma membrane domains in plants
Nature Plants 2024
Jaillais Y, Bayer E, Bergmann D, Botella M, Boutté Y, Bozkurt TO, Caillaud M-C, Germain V, Grossmann G, Heilmann I, Hemsley PA, Kirchhelle C, Martinière A, Miao Y, Mongrand S, Müller S, Noack LC, Oda Y, Ott T, Pan X, Pleskot R, Potocky M, Robert S, Sanchez Rodriguez C, Simon-Plas S, Russinova E, Van Damme D, Van Norman JM, Weijers D, Yalovsky S, Yang Z, Zelazny E, Gronnier J
https://doi.org/10.1038/s41477-024-01742-8
-> See our Nature Plants cover here!
———
Cell wall integrity modulates HOOKLESS1 and PHYTOCHROME INTERACTING FACTOR4 expression controlling apical hook formation
Plant Physiol 2024
Lorrai R, Erguvan Ö, Raggi S, Jonsson K, Široká J, Tarkowská D, Novák O, Griffiths J, Jones AM, Verger S, Robert S, Ferrari S
https://doi.org/10.1093/plphys/kiae370
———
New PEO‑IAA‑inspired anti‑auxins: synthesis, biological activity, and possible application in hemp (Cannabis sativa L.) micropropagation
J Plant Growth Reg 2023, 42:7547-63
Žukauskaitė A, Saiz‑Fernández I, Bieleszová K, Iškauskienė M, Zhang C, Smýkalová I, Dzedulionytė K, Kubeš MF, Sedlářová M, Pařízková B, Pavlović I, Vain T, Petřík I, Malinauskienė V, Šačkus A, Strnad M, Robert S, Napier R, Novák O, Doležal K
https://doi.org/10.1007/s00344-023-11031-x
———
Auxin as an architect of the pectin matrix
J Exp Bot 2023, 74(22):6933-49
Jobert F, Yadav S, Robert S
https://doi.org/10.1093/jxb/erad174
———
Auxin triggers pectin modification during rootlet emergence in white lupin
Plant J 2022, 112(5):1127-40
Jobert F, Soriano A, Brottier L, Casset C, Divol F, Safran J, Lefebvre V, Pelloux J, Robert S*, Péret B* (*joint corresponding authors)
https://doi.org/10.1111/tpj.15993
-> See our Plant Journal cover here!
-> See also the Research Highlight written about our article!
———
Cell biology of the leaf epidermis: Fate specification, morphogenesis and coordination
Plant Cell 2022, 34:209-27
Zuch DT*, Doyle SM*, Majda M, Smith RS, Robert S^, Torii KU^ (* joint first authors; ^joint corresponding authors)
https://doi.org/10.1093/plcell/koab250
———
A network of stress-related genes regulates hypocotyl elongation downstream of selective auxin perception
Plant Physiol 2021, 187(1):430-45
Rigal A*, Doyle SM*, Ritter A, Raggi S, Vain T, O’Brien JA, Goossens A, Pauwels L^, Robert S^ (*joint first authors; ^joint corresponding authors)
https://doi.org/10.1093/plphys/kiab269
-> See our Plant Physiology first author highlight!
———
Solving the puzzle of shape regulation in plant epidermal pavement cells
Ann Rev Plant Biol 2021, 72:525-50
Liu S, Jobert F, Rahneshan Z, Doyle SM*, Robert S* (* joint corresponding authors)
https://doi.org/10.1146/annurev-arplant-080720-081920
———
PICKLE recruits RETINOBLASTOMA RELATED 1 to control lateral root formation in Arabidopsis
Mol Plant Sci 2021, 22:3862
Ötvös K, Miskolczi P, Marhavý P, Cruz-Ramírez A, Benková E, Robert S, Bakó L
https://doi.org/10.3390/ijms22083862
———
The chemical compound ‘Heatin’ stimulates hypocotyl elongation and interferes with the Arabidopsis NIT1-subfamily of nitrilases
Plant J 2021, 106:1523-40
van der Woude L, Piotrowski M, Klaasse G, Paulus JK, Krahn D, Ninck S, Kaschani F, Kaiser M, Novák O, Ljung K, Bulder S, van Verk M, Snoek BL, Fiers M, Martin NI, van der Hoorn RAL, Robert S, Smeekens S, van Zanten M
https://doi.org/10.1111/tjp.15250
———
New fluorescent auxin probes visualize tissue-specific and subcellular distributions of auxin in Arabidopsis
New Phytol 2021, 230:535-49
Pařízková B*, Žukauskaitė A*, Vain T*, Grones P*, Raggi S, Kubeš MF, Kieffer M, Doyle SM, Strnad M, Kepinski S, Napier R, Doležal K, Robert S^, Novák O^ (*joint first authors; ^joint corresponding authors)
https://doi.org/10.1111/nph.17183
———
Cell-surface receptors enable perception of extracellular cytokinins
Nat Comm 2020, 11(1):4284
Antoniadi I, Novák O, Gelová Z, Johnson A, Plíha O, Simerský R, Mik V, Vain T, Mateo-Bonmatí E, Karady M, Pernisová M, Plačková L, Opassathian K, Hejátko J, Robert S, Frim J, Doležal K, Ljung K, Turnbull C
https://doi.org/10.1038/s41467-020-17700-9
———
Fluctuating auxin response gradients determine pavement cell-shape acquisition
Proc Natl Acad Sci USA 2020, 117(27):16027-34
Grones P, Majda M, Doyle SM, Van Damme D, Robert S
https://doi.org/10.1073/pnas.2007400117
———
The CEP5 peptide promotes abiotic stress tolerance, as revealed by quantitative proteomics, and attenuates the AUX/IAA equilibrium in Arabidopsis
Mol & Cell Proteomics 2020, 19(8):1248-62
Smith S, Zhu S, Joos L, Roberts I, Nikonorova N, Vu LD, Stes E, Cho H, Larrieu A, Xuan W, Goodall B, van de Cotte B, Waite JM, Rigal A, Harborough SRR, Persiau G, Vanneste S, Kirschner GK, Vandermarliere E, Martens L, Stahl Y, Audenaert D, Friml J, Felix G, Simon R, Bennett M, Bishopp A, De Jaeger G, Ljung K, Kepinski S, Robert S, Nemhauser J, Hwang I, Gevaert K, Beeckman T, De Smet I
https://doi.org/10.1074/mcp.RA119.001826
———
Polar expedition: mechanisms for protein polar localization
Curr Opin Plant Biol 2020, 53:134-40
Raggi S, Demes E, Liu S, Verger S, Robert S
https://doi.org/10.1016/j.pbi.2019.12.001
———
Auxin: at the crossroads between chemistry and biology
In: The Chemical Biology of Plant Biostimulants (2020), Editors: Geelen D, Xu L
Raggi S, Doyle SM, Robert S
https://doi.org/10.1002/9781119357254.ch5
———
Chemical screening pipeline for identification of specific plant autophagy modulators
Plant Physiol 2019, 181(3):855-66
Dauphinee AN, Cardoso C, Dalman K, Ohlsson JA, Berglund Fick S, Robert S, Hicks GR, Bozhkov P, Minina EA
https://doi.org/10.1104/pp.19.00647
———
A role for the auxin precursor anthranilic acid in root gravitropism via regulation of PIN-FORMED protein polarity and relocalisation in Arabidopsis
New Phytol 2019, 223(3):1420-32
Doyle SM*, Rigal A*, Grones P^, Karady M^, Barange DK, Majda M, Pařízková B, Karampelias M, Zwiewka M, Pencik A, Almqvist F, Ljung K, Novák O, Robert S (*joint first authors; ^joint second authors)
https://doi.org/10.1111/nph.15877
———
Mechanical asymmetry of the cell wall predicts changes in pavement cell geometry
Dev Cell 2019, 50(1):9-10
Majda M, Krupinski P, Jönsson H, Hamant O, Robert S
https://doi.org/10.1016/j.devcel.2019.06.002
———
FORCE-ing the shape
Curr Opin Plant Biol 2019, 52:1–6
Grones P, Raggi S, Robert S
https://doi.org/10.1016/j.pbi.2019.05.008
———
Selective auxin agonists induce specific AUX/IAA protein degradation to modulate plant development
Proc Natl Acad Sci USA 2019, 116(13):6463-72
Vain T*, Raggi S*, Ferro N, Barange DK, Kieffer M, Ma Q, Doyle SM, Thelander M, Pařízková B, Novák O, Ismail A, Enquist PA, Rigal A, Łangowska M, Ramans Harborough S, Zhang Y, Ljung K, Callis J, Almqvist F, Kepinski S, Estelle M, Pauwels L, Robert S (*joint first authors)
https://doi.org/10.1073/pnas.1809037116
———
The inhibitor endosidin 4 targets SEC7 domain-type ARF GTPase exchange factors and interferes with subcellular trafficking in eukaryotes
Plant Cell 2018, 30(10):2553-72
Kania U, Nodzynski T, Lu Q, Hicks GR, Nerinckx W, Mishev K Peurois F, Cherfils J, De Rycke R, Grones P, Robert S, Russinova E, Friml J
https://doi.org/10.1105/tpc.18.00127
———
New fluorescently labeled auxins exhibit promising anti-auxin activity
N Biotechnol 2018, 48:45-52
Bieleszová K, Pařízková B, Kubeš M, Husičková A, Kubala M, Ma Q, Sedlářová M, Robert S, Doležal K, Strnad M, Novák O, Žukauskaitė A
https://doi.org/10.1016/j.nbt.2018.06.003
———
The role of auxin in cell wall expansion
Int J Mol Sci 2018, 19(4):951
Majda M, Robert S
https://doi.org/10.3390/ijms19040951
———
Auxin signaling: a big question to be addressed by small molecules
J Exp Bot 2018, 69(2):313-28
Ma Q, Grones P, Robert S
https://doi.org/10.1093/jxb/erx375
———
Vacuole integrity maintained by DUF300 proteins is required for brassinosteroid signaling regulation
Mol Plant 2017, 11(4):553-67
Liu Q, Vain T*, Viotti C*, Doyle SM*, Tarkowská D, Novák O, Zipfel C, Sitbon F, Robert S, Hofius D (*joint second authors)
https://doi.org/10.1016/j.molp.2017.12.015
———
Mechanochemical polarization of contiguous cell walls shapes plant pavement cells
Dev Cell 2017, 43(3):290–304.e4
Majda M, Grones P, Sintorn I-M, Vain T, Milani P, Krupinski P, Zagórska-Marek B, Viotti C, Jönsson H, Mellerowicz E J, Hamant O, Robert S
https://doi.org/10.1016/j.devcel.2017.10.017
———
Regulating plant physiology with organic electronics
Proc Natl Acad Sci USA 2017, 114(18):4597-602
Poxson DJ, Karady M, Gabrielsson R, Alkattan A Y, Gustavsson A, Doyle SM, Robert S, Ljung K, Grebe M, Simon DT, Berggren M
https://doi.org/10.1073/pnas.1617758114
———
Auxin 2016: a burst of auxin in the warm south of China
Development 2017, 144:533-40
Vernoux T, Robert S
https://doi.org/10.1242/dev.144790
———
2,4-D and IAA amino acid conjugates show distinct metabolism in Arabidopsis
PLoS One 2016, 11(7):e0159269
Eyer L, Vain T, Pařízková B, Oklestkova J, Barbez E, Kozubíková H, Pospíšil T, Wierzbicka R, Kleine-Vehn J, Fránek M, Strnad M, Robert S*, Novak O* (*joint corresponding authors)
https://doi.org/10.1371/journal.pone.0159269
———
Mitochondrial uncouplers inhibit clathrin-mediated endocytosis largely through cytoplasmic acidification
Nat Comm 2016, 7:11710
Dejonghe W, Kuenen S, Mylle E, Vasileva M, Keech O, Viotti C, Swerts J, Fendrych M, Ortiz-Morea FA, Mishev K, Delang S, Scholl S, Zarza X, Heilmann M, Kourelis J, Kasprowicz J, Nguyen LSL, Drozdzecki A, Van Houtte I, Szatmári A-M, Majda M, Baisa G, Bednarek SY, Robert S, Audenaert D, Testerink C, Munnik T, Van Damme D, Heilmann I, Schumacher K, Winne J, Friml J, Verstreken P, Russinova E
https://doi.org/10.1038/ncomms11710
———
Extra- and intracellular distribution of cytokinins in the leaves of monocots and dicots
N Biotechnol 2016, 33(5):735-42
Jiskrová E, Novák O, Pospíšilová H, Holubová K, Karády M, Galuszka P, Robert S, Frébort I
https://doi.org/10.1016/j.nbt.2015.12.010
———
Small molecules unravel complex interplay between auxin biology and endomembrane trafficking
J Exp Bot 2015, 66(16):4971-82
Doyle SM, Vain T, Robert S
https://doi.org/10.1093/jxb/erv179
———
Osmotic stress modulates the balance between exocytosis and clathrin-mediated endocytosis in Arabidopsis thaliana
Mol Plant 2015, 8(8):1175-87
Zwiewka M, Nodzyński T, Robert S, Vanneste S, Friml J
https://doi.org/10.1016/j.molp.2015.03.007
———
An early secretory pathway mediated by GNOM-LIKE 1 and GNOM is essential for basal polarity establishment in Arabidopsis thaliana
Proc Natl Acad Sci USA 2015, 112(7):E806-15
Doyle SM, Haeger A, Vain T, Rigal A, Viotti C, Łangowska M, Ma Q, Friml J, Raikhel NV, Hicks GR, Robert S
https://doi.org/10.1073/pnas.1424856112
———
Live cell imaging of FM4-64, a tool for tracing the endocytic pathways in Arabidopsis root cells
Methods Mol Biol 2015, 1242:93-103
Rigal A, Doyle SM, Robert S
https://doi.org/10.1007/978-1-4939-1902-4_9
———
Unraveling plant hormone signaling through the use of small molecules
Front Plant Sci 2014, 5:373
Rigal A, Ma Q, Robert S
https://doi.org/10.3389/fpls.2014.00373
———
The cellulase KORRIGAN is part of the cellulose synthase complex
Plant Physiol 2014, 165:1521-32
Vain T, Crowell EF, Timpano H, Biot E, Desprez T, Mansoori N, Trindade LM, Pagant S, Robert S, Höfte H, Gonneau M, Vernhettes S
https://doi.org/10.1104/pp.114.241216
———
Trafficking modulator TENin1 inhibits endocytosis, causes endomembrane protein accumulation at the pre-vacuolar compartment and impairs gravitropic response in Arabidopsis thaliana
Biochem J 2014, 460:177-85
Paudyal R, Jamaluddin A, Warren JP, Doyle SM, Robert S, Warriner SL, Baker A
https://doi.org/10.1042/BJ20131136
———
Using a reverse genetics approach to investigate small-molecule activity
Methods Mol Biol 2014, 1056:51-62
Doyle SM, Robert S
https://doi.org/10.1007/978-1-62703-592-7_6
———
Auxin biology revealed by small molecules
Physiol Plant 2014, 151(1):25-42
Ma Q, Robert S
https://doi.org/10.1111/ppl.12128
———
The use of chemical biology to study plant cellular processes – subcellular trafficking
In: Plant Chemical Biology (2013), Editors: Audenaert D, Overvoorde P
Haeger A, Łangowska M, Robert S
https://b-ok.cc/book/2357338/d7383e
———
ABCG9, ABCG11 and ABCG14 ABC transporters are required for vascular development in Arabidopsis
Plant J 2013, 76(5):811-24
Le Hir R, Sorin C, Chakraborti D, Moritz T, Schaller H, Tellier F, Robert S, Morin H, Bako L, Bellini C
https://doi.org/10.1111/tpj.12334
———
ECHIDNA-mediated post-Golgi trafficking of auxin carriers for differential cell elongation
Proc Natl Acad Sci USA 2013, 110(40):16259-64
Boutté Y, Jonsson K, McFarlane HE, Johnson E, Gendre D, Swarup R, Friml J, Samuels L, Robert S, Bhalerao RP
https://doi.org/10.1073/pnas.1309057110
———
Defining the selectivity of processes along the auxin response chain: a study using auxin analogues
New Phytol 2013, 200(4):1034-48
Simon S, Kubeš M, Baster P, Robert S, Dobrev PI, Friml J, Petrášek J, Zažímalová E
https://doi.org/10.1111/nph.12437
———
The caspase-related protease separase (extra spindle poles) regulates cell polarity and cytokinesis in Arabidopsis
Plant Cell 2013, 25(6):2171-86
Moschou PN, Smertenko AP, Minina EA, Fukada K, Savenkov EI, Robert S, Hussey PJ, Bozhkov PV
https://doi.org/10.1105/tpc.113.113043
———
Cell polarity and patterning by PIN trafficking through early endosomal compartments in Arabidopsis thaliana
PLoS Genet 2013, 9(5):e1003540
Tanaka H, Kitakura S, Rakusová H, Uemura T, Feraru MI, De Rycke R, Robert S, Kakimoto T, Friml J
https://doi.org/10.1371/journal.pgen.1003540
———
Auxin: simply complicated
J Exp Bot 2013, 64(9):2565-77
Sauer M, Robert S, Kleine-Vehn J
https://doi.org/10.1093/jxb/ert139
———
ROOT ULTRAVIOLET B-SENSITIVE1/WEAK AUXIN RESPONSE3 is essential for polar auxin transport in Arabidopsis
Plant Physiol 2013, 162(2):965-76
Yu H, Karampelias M, Robert S, Peer WA, Swarup R, Ye SQ, Ge L, Cohen J, Murphy A, Friml J, Estelle M
https://doi.org/10.1104/pp.113.217018
———
SCF(TIR1/AFB)-auxin signalling regulates PIN vacuolar trafficking and auxin fluxes during root gravitropism
EMBO J 2012, 32(2):260-74
Baster P, Robert S, Kleine-Vehn J, Vanneste S, Kania U, Grunewald W, De Rybel B, Beeckman T, Friml J
https://doi.org/10.1038/emboj.2012.310
———
ABP1 and ROP6 GTPase signaling regulate clathrin-mediated endocytosis in Arabidopsis roots
Curr Biol 2012, 22(14):1326-32
Chen X, Naramoto S, Robert S, Tejos R, Löfke C, Lin D, Yang Z, Friml J
https://doi.org/10.1016/j.cub.2012.05.020
———
Recycling, clustering, and endocytosis jointly maintain PIN auxin carrier polarity at the plasma membrane
Mol Syst Biol 2011, 7:540
Kleine-Vehn J, Wabnik K, Martiniere A, Langowski L, Willig K, Naramoto S, Leitner J, Tanaka H, Jakobs S, Robert S, Luschnig C, Govaerts W, Hell SW, Runions J, Friml J
https://doi.org/10.1038/msb.2011.72
———
Clusters of bioactive compounds target dynamic endomembrane networks in vivo
Proc Natl Acad Sci USA 2011, 108(43):17850-5
Drakakaki G*, Robert S*, Szatmari A*, Brown MQ, Nagawa S, Van Damme D, Leonard M, Yang Y, Girke T, Schmid SL, Russinova E, Friml J, Raikhel NV, Hicks GR (*joint first authors)
https://doi.org/10.1073/pnas.1108581108
———
Monoubiquitin-dependent endocytosis of the IRON-REGULATED TRANSPORTER 1 (IRT1) transporter controls iron uptake in plants
Proc Natl Acad Sci USA 2011, 108(32):E450-8
Barberon M, Zelazny E, Robert S, Conéjéro G, Curie C, Friml J, Vert G
https://doi.org/10.1073/pnas.1100659108
———
Clathrin mediates endocytosis and polar distribution of PIN auxin transporters in Arabidopsis
Plant Cell 2011, 23:1920-31
Kitakura S, Vanneste S, Robert S, Löfke C, Teichmann T, Tanaka H, Friml
https://doi.org/10.1105/tpc.111.083030
———
ADP-ribosylation factor machinery mediates endocytosis in plant cells
Proc Natl Acad Sci USA 2010, 107:21890-5
Naramoto S, Kleine-Vehn J, Robert S, Fujimoto M, Dainobu T, Paciorek T, Ueda T, Nakano A, Van Montagu MCE, Fukuda H, Friml J
https://doi.org/10.1073/pnas.1016260107
———
ABP1 mediates auxin inhibition of clathrin-dependent endocytosis in Arabidopsis
Cell 2010, 143(1):111-21
https://doi.org/10.1o16/j.cell.2010.09.027
———
Chemical dissection of endosomal pathways
Plant Signal Behav 2009, 4(1):57-62
https://doi.org/10.4161/psb.4.1.7314
———
Powerful partners: Arabidopsis and chemical genomics
https://doi.org/10.1199/tab.0109
Nat Protoc 2007, 2(2):259-62
Robert S, Zouhar J, Carter C, Raikhel N
https://doi.org/10.1038/nprot.2007.26
———
Divergent functions of VTI12 and VTI11 in trafficking to storage and lytic vacuoles in Arabidopsis
Proc Natl Acad Sci USA 2007, 104(9):3645-50
https://doi.org/10.1073/pnas.0611147104
———
Arabidopsis reversibly glycosylated polypeptides 1 and 2 are essential for pollen development
https://doi.org/10.1104/pp.106.086363
———
An Arabidopsis endo-1,4-beta-D-glucanase involved in cellulose synthesis undergoes regulated intracellular cycling
https://doi.org/10.1105/tpc.105.036228
———